Characterizing Dietary Niche Segregation at High Resolution
An open access article published in Ecology & Evolution details a new high resolution method for niche characterization using carbon isotope patterns of essential amino acids
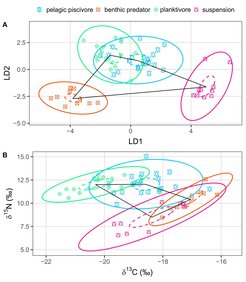
Niche segregation of consumers from the Baltic Sea demonstrated with two different analytical approaches. Stable isotope fingerprinting of amino acids (a) can resolve dietary niches at a much higher resolution than bulk stable isotope analysis (b).
In nature, niche partitioning helps species to reduce competitive interactions and facilitates efficient and effective utilization of resources. Although niche partitioning has been examined extensively, it has been notoriously difficult to characterize resource use at high resolution with prevailing methods. While biomarkers such as bulk stable isotopes have proven useful in many instances for illuminating trophic niches, they may lack in resolution, especially when spatiotemporal variability in a system is high. This methodological gap has made it difficult to model and understand complex life systems, particularly marine food webs, because they tend to be highly compartmentalized.
A new open access research paper published in Ecology & Evolution demonstrates how a novel method, carbon isotope (d13C) patterns of essential amino acids (EAAs), also termed d13CEAA fingerprints, can characterize niche differentiation in a highly dynamic marine system. By examining 10 different species from the Baltic Sea, a regional sea with strong salinity and temperature gradients, the study reveals that the d13CEAA fingerprints can differentiate among the trophic niches of both functional groups and species at an unprecedented resolution. The differences that the new method is able to measure are likely driven by how consumers rely on particular energy channels, i.e. in phytoplankton assemblages, that each have distinct fingerprints. Since these fingerprints remain largely invariant across different environmental conditions, they can be leveraged across time and space to characterize niche partitions. The study found that bulk isotopes were considerably less informative of dietary niches than d13CEAA fingerprints.
The methodological advance presented in this study can improve our understanding of how environmental stressors and management practices affect the functions of a marine ecosystem. Current marine food webs are predicted to be fragile and susceptible to structural changes with consequent alterations in the functioning of the ecosystem. As environmental changes are accelerating, it is crucial to understand whether and how quickly marine food webs can adapt to changes in phytoplankton assemblages and top predator abundances. For this reason, identifying and quantifying feeding interactions across trophic levels, from phytoplankton to zooplankton to higher trophic levels, is key.
