Recombination-aware reconstruction of Hepatitis B virus phylogenies
Hundreds of millions of people from all continents are currently infected with Hepatitis B virus (HBV). HBV is transmitted through close contact with infected body fluids, including sexual and perinatal transmission, and has no known environmental or animal reservoir. Its dissemination is therefore tightly linked to the movement of humans, whose past population dynamics and migration routes have likely largely shaped the current structure of its genetic diversity. Despite dramatic advances in molecular virology during the last decades, and the discovery of ancient HBV genomes from archaeological remains, much mystery persists regarding the evolutionary history of this virus.
HBV is known to undergo frequent genetic recombination. This leads to horizontal transfers of genetic material between strains and can introduce strong biases in phylogenetic reconstructions that only consider vertical diversification processes. Accurate representation of the evolutionary history of this virus would therefore require the use of phylogenetic models explicitly accounting for genetic recombination occurring between lineages, which is known to be a challenging mathematical and computational problem.
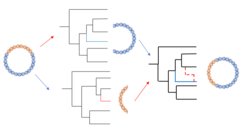
With this project, we aim to build upon an existing method that was developed for the analysis of recombining bacterial genomes (the clonalOrigin model implemented in the BEAST2 package Bacter) for the analysis of HBV molecular datasets. This includes the adaptation of the method to short circular genomes, tuning of the MCMC sampling to improve inference, and refinement of the posterior probability representation of estimated ancestral recombination graphs. Ultimately, we will apply the method to a dataset comprising modern and ancient HBV genome sequences in order to get further insights into the evolutionary history of this virus.
